Bootstrap Boosting
bboost.RdWrapper function for applying bootstrap estimation using gradient boosting.
## Bootstrap boosting.
bboost(..., data, type = 1, cores = 1,
n = 2, prob = 0.623, fmstop = NULL,
trace = TRUE, drop = FALSE, replace = FALSE)
## Plotting function.
bboost_plot(object, col = NULL)
## Predict method.
# S3 method for bboost
predict(object, newdata, ..., cores = 1, pfun = NULL)Arguments
- ...
Arguments passed to
bamlssandpredict.bamlss.- data
The data frame to be used for modeling.
- type
Type of algorithm,
type = 1uses all observations and samples with replacement,type = 2uses only a fraction specified inproband samples with replacement.- cores
The number of cores to be used.
- n
The number of bootstrap iterations.
- prob
The fraction that should be used to fit the model in each bootstrap iteration.
- fmstop
The function that should return the optimum stopping iteration. The function must have two arguments: (1) the
modelend (2) thedata. The function must return a list with two named arguments: (1)"mstop"the optimum stopping iteration and (2) a vector of the objective criterion that should be evaluated by the hold out sample data during each bootstrap iteration. See the examples.- trace
Prints out the current state of the bootstrap algorithm.
- drop
Should only the best set of parameters be saved?
- replace
Sampling with replacement, or sampling
ceiling(nobs * prob)rows of the data for fitting thenmodels.- object
The
"bboost"object used for prediction and plotting.- col
The color that should be used for plotting.
- newdata
The data frame predictions should be made for.
- pfun
The prediction function that should be used, for example
predictncould be used, too. Note that this is experimental.
Value
A list of bamlss objects.
Examples
if (FALSE) ## Simulate data.
set.seed(123)
d <- GAMart()
## Estimate model.
f <- num ~ s(x1) + s(x2) + s(x3) + s(lon,lat)
## Function for evaluation of hold out sample
## criterion to find the optimum mstop.
fmstop <- function(model, data) {
p <- predict(model, newdata = data, model = "mu")
mse <- NULL
for(i in 1:nrow(model$parameters))
mse <- c(mse, mean((data$num - p[, i])^2))
list("MSE" = mse, "mstop" = which.min(mse))
}
## Bootstrap boosted models.
b <- bboost(f, data = d, n = 50, cores = 3, fmstop = fmstop)
## Plot hold out sample MSE.
bboost_plot(b)
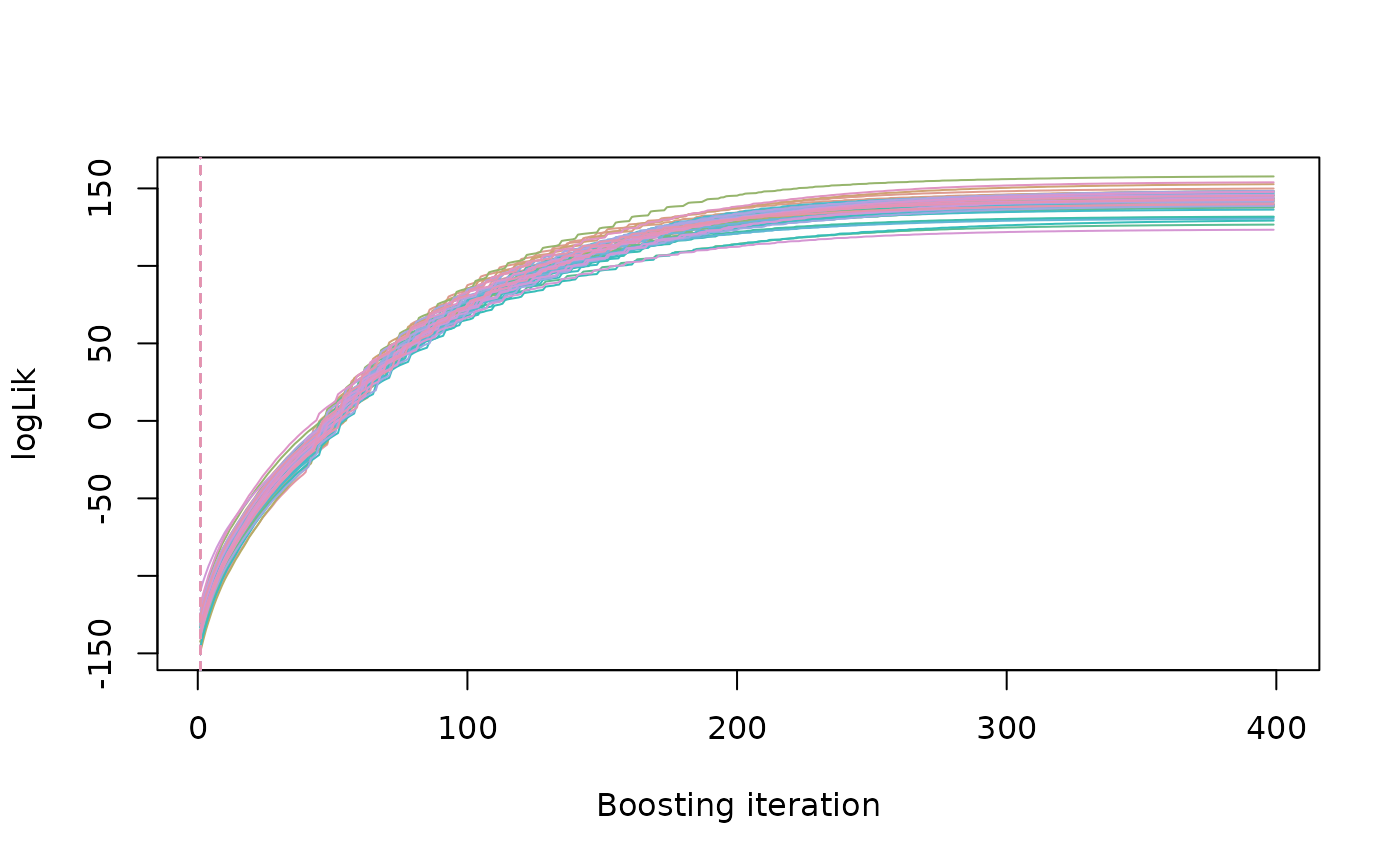 ## Predict for each bootstrap sample.
nd <- data.frame("x2" = seq(0, 1, length = 100))
p <- predict(b, newdata = nd, model = "mu", term = "x2")
plot2d(p ~ x2, data = nd)
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'as.matrix': object 'p' not found
## Predict for each bootstrap sample.
nd <- data.frame("x2" = seq(0, 1, length = 100))
p <- predict(b, newdata = nd, model = "mu", term = "x2")
plot2d(p ~ x2, data = nd)
#> Error in h(simpleError(msg, call)): error in evaluating the argument 'x' in selecting a method for function 'as.matrix': object 'p' not found